Various utils
Flowpaths implements various helper functions on graphs. They can be access with the prefix flowpaths.utils.
Graph visualization and drawing
You can create drawing as this one
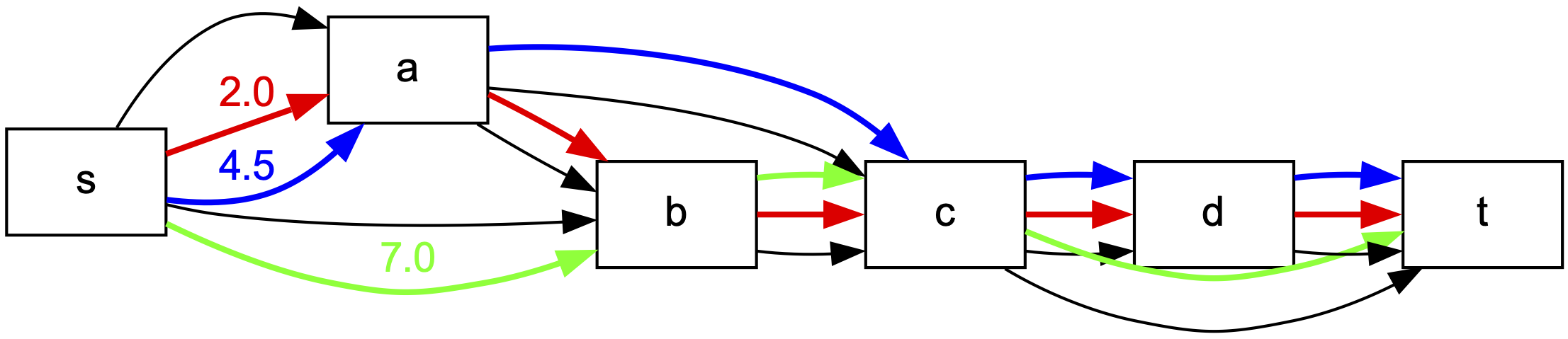
using the following code:
import flowpaths as fp
import networkx as nx
# Create a simple graph
graph = nx.DiGraph()
graph.graph["id"] = "simple_graph"
graph.add_edge("s", "a", flow=6)
graph.add_edge("s", "b", flow=7)
graph.add_edge("a", "b", flow=2)
graph.add_edge("a", "c", flow=5)
graph.add_edge("b", "c", flow=9)
graph.add_edge("c", "d", flow=6)
graph.add_edge("c", "t", flow=7)
graph.add_edge("d", "t", flow=6)
# Solve the minimum path error model
mpe_model = fp.kMinPathError(graph, flow_attr="flow", k=3, weight_type=float)
mpe_model.solve()
# Draw the solution
if mpe_model.is_solved():
solution = mpe_model.get_solution()
fp.utils.draw(
G=graph,
filename="simple_graph.pdf",
flow_attr="flow",
paths=solution["paths"],
weights=solution["weights"],
draw_options={
"show_graph_edges": True,
"show_edge_weights": False,
"show_path_weights": False,
"show_path_weight_on_first_edge": True,
"pathwidth": 2,
})
This produces a file with extension .pdf storing the PDF image of the graph.
Sankey Diagram Visualization
For acyclic graphs (DAGs), you can create interactive Sankey diagrams using plotly. Sankey diagrams are particularly effective for visualizing flow decompositions, as they show:
- Each node in the graph as a labeled box
- Each path as a colored flow whose width represents the path weight
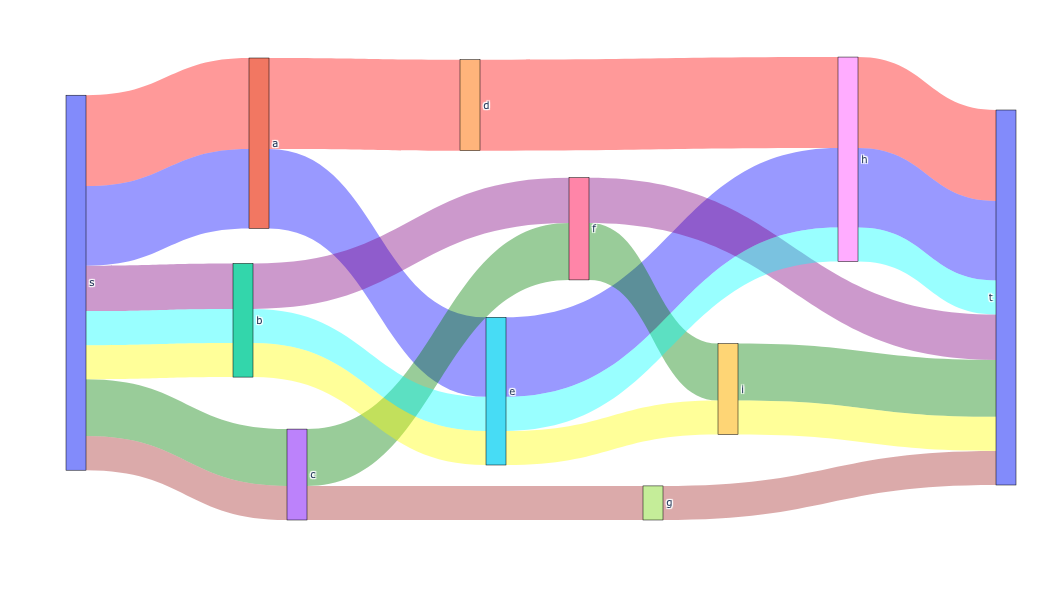
To create a Sankey diagram, set "style": "sankey" in the draw_options:
import flowpaths as fp
import networkx as nx
# Create a sample DAG
G = nx.DiGraph()
G.add_edge('s', 'a', flow=10)
G.add_edge('s', 'b', flow=5)
G.add_edge('a', 'c', flow=6)
G.add_edge('a', 'd', flow=4)
G.add_edge('b', 'c', flow=3)
G.add_edge('b', 'd', flow=2)
G.add_edge('c', 't', flow=9)
G.add_edge('d', 't', flow=6)
# Compute minimum flow decomposition
solver = fp.MinFlowDecomp(G, flow_attr='flow')
solver.solve()
solution = solver.get_solution()
# Draw as interactive Sankey diagram
fp.utils.draw(
G=G,
filename="flow_sankey.html", # saves as HTML (interactive)
flow_attr='flow',
paths=solution['paths'],
weights=solution['weights'],
draw_options={
"style": "sankey"
}
)
Features:
- Interactive: Hover over nodes and links to see details, zoom and pan the diagram
- Jupyter support: Automatically displays inline when run in Jupyter notebooks
- Dual output: Automatically saves both HTML (interactive) and a static image (PDF by default)
- Automatic coloring: Each path gets a distinct color; shared edges show blended colors
- Graph identification: Uses the graph’s ID as the diagram title if available
Requirements:
- plotly: Installed automatically with flowpaths
- kaleido: Installed automatically with flowpaths for static image export
File formats:
The function automatically saves both formats:
- HTML file (interactive): [basename].html
- Static image: [basename].pdf (or .png, .svg if specified)
# Saves both output.html and output.pdf
fp.utils.draw(G, "output", paths=paths, weights=weights,
draw_options={"style": "sankey"})
# Saves both flow.html and flow.png
fp.utils.draw(G, "flow.png", paths=paths, weights=weights,
draw_options={"style": "sankey"})
# Saves both diagram.html and diagram.svg
fp.utils.draw(G, "diagram.svg", paths=paths, weights=weights,
draw_options={"style": "sankey"})
Note: Sankey diagrams require the graph to be acyclic (DAG). If the graph contains cycles, use the traditional graphviz rendering ("style": "default" or "style": "points").
See examples/sankey_demo.py and examples/sankey_demo.ipynb for complete examples.
Logging
flowpaths exposes a simple logging helper via fp.utils.configure_logging. Use it to control verbosity, enable console/file logging, and set file mode.
Basic usage (console logging at INFO level):
import flowpaths as fp
fp.utils.configure_logging(
level=fp.utils.logging.INFO,
log_to_console=True,
)
Also log to a file (append mode):
fp.utils.configure_logging(
level=fp.utils.logging.DEBUG, # default is DEBUG
log_to_console=True, # show logs in terminal
log_file="flowpaths.log", # write logs to this file
file_mode="a", # "a" append (or "w" overwrite)
)
Notes:
- Levels available: fp.utils.logging.DEBUG, INFO, WARNING, ERROR, CRITICAL.
- Default level is DEBUG. If you prefer quieter output, use INFO or WARNING.
- Internally, the package logs through its own logger; configure_logging sets handlers/formatters accordingly.
API reference:
Configures logging for the flowpaths package.
Parameters:
-
level: int, optionalLogging level (e.g., fp.utils.logging.DEBUG, fp.utils.logging.INFO). Default is fp.utils.logging.DEBUG.
-
log_to_console: bool, optionalWhether to log to the console. Default is True.
-
log_file: str, optionalFile path to log to. If None, logging to a file is disabled. Default is None. If a file path is provided, the log will be written to that file. If the file already exists, it will be overwritten unless
file_modeis set to “a”. -
file_mode: str, optionalMode for the log file. “a” (append) or “w” (overwrite). Default is “w”.
Source code in flowpaths/utils/logging.py
check_flow_conservation
Check if the flow conservation property holds for the given graph.
Parameters
-
G: nx.DiGraphThe input directed acyclic graph, as networkx DiGraph.
-
flow_attr: strThe attribute name from where to get the flow values on the edges.
Returns
-
bool:
True if the flow conservation property holds, False otherwise.
Source code in flowpaths/utils/graphutils.py
draw
draw(
G: DiGraph,
filename: str,
flow_attr: str = None,
paths: list = [],
weights: list = [],
additional_starts: list = [],
additional_ends: list = [],
subpath_constraints: list = [],
draw_options: dict = {
"show_graph_edges": True,
"show_edge_weights": False,
"show_node_weights": False,
"show_graph_title": False,
"show_path_weights": False,
"show_path_weight_on_first_edge": True,
"pathwidth": 3.0,
"style": "default",
"color_nodes": False,
"sankey_arrowlen": 0,
"sankey_color_toggle": False,
"sankey_arrow_toggle": False,
},
)
Draw the graph with the paths and their weights highlighted.
Parameters
-
G: nx.DiGraphThe input directed acyclic graph, as networkx DiGraph.
-
filename: strThe name of the file to save the drawing. The file type is inferred from the extension. Supported extensions are ‘.bmp’, ‘.canon’, ‘.cgimage’, ‘.cmap’, ‘.cmapx’, ‘.cmapx_np’, ‘.dot’, ‘.dot_json’, ‘.eps’, ‘.exr’, ‘.fig’, ‘.gd’, ‘.gd2’, ‘.gif’, ‘.gtk’, ‘.gv’, ‘.ico’, ‘.imap’, ‘.imap_np’, ‘.ismap’, ‘.jp2’, ‘.jpe’, ‘.jpeg’, ‘.jpg’, ‘.json’, ‘.json0’, ‘.pct’, ‘.pdf’, ‘.pic’, ‘.pict’, ‘.plain’, ‘.plain-ext’, ‘.png’, ‘.pov’, ‘.ps’, ‘.ps2’, ‘.psd’, ‘.sgi’, ‘.svg’, ‘.svgz’, ‘.tga’, ‘.tif’, ‘.tiff’, ‘.tk’, ‘.vml’, ‘.vmlz’, ‘.vrml’, ‘.wbmp’, ‘.webp’, ‘.x11’, ‘.xdot’, ‘.xdot1.2’, ‘.xdot1.4’, ‘.xdot_json’, ‘.xlib’
-
flow_attr: strThe attribute name from where to get the flow values on the edges. Default is an empty string, in which case no edge weights are shown.
-
paths: listThe list of paths to highlight, as lists of nodes. Default is an empty list, in which case no path is drawn. Default is an empty list.
-
weights: listThe list of weights corresponding to the paths, of various colors. Default is an empty list, in which case no path is drawn.
-
additional_starts: listA list of additional nodes to highlight in green as starting nodes. Default is an empty list. -
additional_ends: listA list of additional nodes to highlight in red as ending nodes. Default is an empty list. -
subpath_constraints: listA list of subpaths to highlight in the graph as dashed edges, of various colors. Each subpath is a list of edges. Default is an empty list. There is no association between the subpath colors and the path colors.
-
draw_options: dictA dictionary with the following keys:
-
show_graph_edges: boolWhether to show the edges of the graph. Default is
True. -
show_edge_weights: boolWhether to show the edge weights in the graph from the
flow_attr. Default isFalse. -
show_node_weights: boolWhether to show the node weights in the graph from the
flow_attr. Default isFalse. -
show_graph_title: boolWhether to show the graph title (from graph id) in the figure. Default is
False. -
show_path_weights: boolWhether to show the path weights in the graph on every edge. Default is
False. -
show_path_weight_on_first_edge: boolWhether to show the path weight on the first edge of the path. Default is
True. -
pathwidth: floatThe width of the path to be drawn. Default is
3.0. -
style: strThe style of the drawing. Available options:
default,points,sankey.default: Standard graphviz rendering with nodes as rounded rectanglespoints: Graphviz rendering with nodes as pointssankey: Interactive Sankey diagram using plotly (requires acyclic graph). Saves as HTML by default (interactive) or static image formats (png, pdf, svg) if kaleido is installed. Automatically displays in Jupyter notebooks.
-
color_nodes: boolWhether to use the existing node coloring behavior. If `False` (default), all nodes use a neutral color. If `True`, nodes are colored as before (including `additional_starts` in green and `additional_ends` in red for graphviz styles). -
sankey_arrowlen: floatLength of arrowheads for Sankey links (Plotly
arrowlen). Default is0(no arrowheads). -
sankey_color_toggle: boolWhether to add an interactive toggle (buttons) to switch Sankey links between colored and monochrome gray. Default is
False. -
sankey_arrow_toggle: boolWhether to add an interactive toggle (buttons) to switch Sankey link arrowheads on/off. Default is
False.
-
Source code in flowpaths/utils/graphutils.py
551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 | |
fpid
get_subgraph_between_topological_nodes
get_subgraph_between_topological_nodes(
graph: DiGraph,
topo_order: list,
left: int,
right: int,
) -> nx.DiGraph
Create a subgraph with the nodes between left and right in the topological order, including the edges between them, but also the edges from these nodes that are incident to nodes outside this range.
Source code in flowpaths/utils/graphutils.py
max_bottleneck_path
Computes the maximum bottleneck path in a directed graph.
Parameters
-
G: nx.DiGraphA directed graph where each edge has a flow attribute.
-
flow_attr: strThe flow attribute from where to get the flow values.
Returns
-
tuple: A tuple containing:
- The value of the maximum bottleneck.
- The path corresponding to the maximum bottleneck (list of nodes). If no s-t flow exists in the network, returns (None, None).
Source code in flowpaths/utils/graphutils.py
max_occurrence
Check what is the maximum number of edges of seq that appear in some path in the list paths_in_DAG.
This assumes paths_in_DAG are paths in a directed acyclic graph.
Parameters
- seq (list): The sequence of edges to check.
- paths (list): The list of paths to check against, as lists of nodes.
Returns
- int: the largest number of seq edges that appear in some path in paths_in_DAG
Source code in flowpaths/utils/graphutils.py
read_graph
Parse a single graph block from a list of lines.
Accepts one or more header lines at the beginning (each prefixed by ‘#’), followed by a line containing the number of vertices (n), then any number of edge lines of the form: “u v w” (whitespace-separated).
Subpath constraint lines
Lines starting with “#S” define a (directed) subpath constraint as a sequence of nodes: “#S n1 n2 n3 …”. For each such line we build the list of consecutive edge tuples [(n1,n2), (n2,n3), …] and append this edge-list (the subpath) to G.graph[“constraints”]. Duplicate filtering is applied on the whole node sequence: if an identical sequence of nodes has already appeared in a previous “#S” line, the entire subpath line is ignored (its edges are not added again). Different subpaths may
share edges; they are kept as separate entries. After all graph edges are parsed, every constraint edge is validated to ensure it exists in the graph; a missing edge raises ValueError.
Example block
graph number = 1 name = foo
any other header line
S a b c d (adds subpath [(a,b),(b,c),(c,d)])
S b c e (adds subpath [(b,c),(c,e)])
S a b c d (ignored: exact node sequence already seen)
5 a b 1.0 b c 2.5 c d 3.0 c e 4.0
Source code in flowpaths/utils/graphutils.py
20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 | |
read_graphs
Read one or more graphs from a file.
Supports graphs whose header consists of one or multiple consecutive lines prefixed by ‘#’. Each graph block is: - one or more header lines starting with ‘#’ - one line with the number of vertices (n) - zero or more edge lines “u v w”
Graphs are delimited by the start of the next header (a line starting with ‘#’) or the end of file.
Source code in flowpaths/utils/graphutils.py
read_ngraph
Parse a single node-weighted ngraph block from a list of lines.
Expected block structure
- one or more leading header lines starting with ‘#’ (optional #S constraints can appear here)
- one line with the number of nodes n
- a marker line starting with ‘#NODES’
- exactly n node lines: “node_id node_weight”
- a marker line starting with ‘#EDGES’
- zero or more edge lines: “u v edge_weight”
Constraint lines
- ‘#S n1 n2 n3 …’ lines define subpath constraints.
- Duplicates are filtered by exact node sequence.
- Constraints are stored in G.graph[‘constraints’] as edge lists.
Source code in flowpaths/utils/graphutils.py
185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 | |
read_ngraphs
Read one or more ngraph blocks from a file.
Graph blocks are delimited by lines starting with ‘# graph’ (case-insensitive). If no such delimiter exists, the whole file is parsed as one ngraph block.